
AlphaFold vs ESMFold: Complete Comparison of AI Protein Structure Prediction Models
A detailed comparison of AlphaFold vs ESMFold, exploring architecture, accuracy, speed, computational cost, real-world applications, and future impact in AI-driven protein structure prediction.
MODERN DISEASESHEALTH/DISEASEAI/FUTURECOMPANY/INDUSTRY
Sachin K Chaurasiya
3/6/20265 min read


Protein structure determines biological function. For decades, predicting 3D protein shapes from amino acid sequences was one of biology’s hardest problems. That changed with the rise of deep learning models like AlphaFold and ESMFold.
Both systems predict protein structures with remarkable accuracy, but they are built on very different philosophies. This article explores their architecture, performance, scalability, technical foundations, limitations, and future implications in depth.
What Is AlphaFold?
AlphaFold is an AI system developed by DeepMind, part of Google. It achieved a breakthrough at the CASP14 competition in 2020 by predicting protein structures with near-experimental accuracy.
Technical Foundation
AlphaFold 2 introduced several innovations:
Evoformer architecture for integrating sequence and structural reasoning
Attention mechanisms across multiple sequence alignments (MSA)
Iterative refinement cycles that progressively improve structure
End-to-end differentiable training
It combines:
Evolutionary data
Structural templates
Geometric reasoning
Confidence estimation metrics like pLDDT
AlphaFold Database Impact
DeepMind released the AlphaFold Protein Structure Database containing hundreds of millions of predicted structures. This has:
Accelerated biological research worldwide
Reduced dependency on X-ray crystallography and cryo-EM for initial exploration
Enabled drug target discovery in record time
What Is ESMFold?
ESMFold was developed by Meta AI. It is based on large-scale protein language models trained on massive protein sequence datasets.
Core Innovation
Instead of relying on evolutionary alignments like AlphaFold, ESMFold:
Uses transformer-based protein language models
Learns structural patterns directly from sequence data
Predicts 3D structures from single sequences
Eliminates the need for computationally expensive MSAs
This makes ESMFold significantly faster and more scalable.

Deep Technical Comparison
Data Dependency
AlphaFold
Requires large evolutionary databases
Uses MSAs extensively
Performance improves with rich homologous sequence data
ESMFold
Trained on billions of protein sequences
Does not require MSA during inference
Works better when evolutionary information is sparse
Key Insight: AlphaFold extracts evolutionary signal explicitly. ESMFold internalizes it during pretraining.
Model Architecture
AlphaFold
Evoformer blocks
Pair representation modules
Structure module with geometric constraints
Recycling mechanism for iterative refinement
ESMFold
Large transformer encoder (protein language model)
Attention layers across amino acid tokens
Lightweight structure prediction head
Single forward pass structure inference
AlphaFold focuses on precision and refinement. ESMFold focuses on efficient generalization.
Computational Efficiency
AlphaFold
Heavy MSA generation step
GPU-intensive inference
Longer processing time per protein
ESMFold
No MSA pipeline
Faster runtime
Lower GPU and memory requirements
This makes ESMFold suitable for large-scale metagenomic projects.
Training Paradigm
AlphaFold
Supervised learning on protein structure databases
Incorporates structural labels from experimental data
ESMFold
Self-supervised pretraining on sequence data
Structure prediction added after language model training
This difference reflects two AI philosophies:
Structure-first modeling (AlphaFold)
Language-first modeling (ESMFold)


Computational Efficiency
AlphaFold
Heavy MSA generation step
GPU-intensive inference
Longer processing time per protein
ESMFold
No MSA pipeline
Faster runtime
Lower GPU and memory requirements
This makes ESMFold suitable for large-scale metagenomic projects.
Training Paradigm
AlphaFold
Supervised learning on protein structure databases
Incorporates structural labels from experimental data
ESMFold
Self-supervised pretraining on sequence data
Structure prediction added after language model training
This difference reflects two AI philosophies:
Structure-first modeling (AlphaFold)
Language-first modeling (ESMFold)
Real-World Applications
Drug Discovery
AlphaFold is widely used in:
Target structure identification
Binding site prediction
Molecular docking preparation
ESMFold is used for:
Rapid candidate screening
Identifying promising structural leads
Metagenomics and Microbial Research
ESMFold shines in:
Massive environmental protein datasets
Novel organisms
Rapid annotation pipelines
AlphaFold works well but requires heavier preprocessing.
Protein Engineering
Both models support:
Enzyme optimization
Stability prediction
Mutational impact studies
AlphaFold may offer slightly better refinement for engineered proteins.
Academic Research
AlphaFold:
Preferred for publication-grade structural analysis
Strong confidence scoring metrics
ESMFold:
Ideal for exploratory hypothesis generation
Confidence Metrics and Reliability
AlphaFold
Provides:
pLDDT score (per-residue confidence)
Predicted aligned error (PAE)
These metrics help researchers assess reliability.
ESMFold
Also provides confidence scores, but:
Generally less detailed than AlphaFold
Slightly less interpretability in structural uncertainty
Limitations of Both Models
Despite their power, neither system fully replaces experimental methods.
Common Challenges
Predicting dynamic conformations
Modeling protein-protein interactions
Accounting for post-translational modifications
Predicting ligand-bound states
Handling intrinsically disordered proteins
Protein folding in living cells is influenced by environment, chaperones, and biochemical context. AI models approximate but do not replicate this complexity.
AlphaFold vs ESMFold for AI and Biotech Startups
If you are building tools in biotech or computational biology:
Use ESMFold for scalable APIs and rapid prediction services
Use AlphaFold when structural precision directly impacts downstream modeling
Many modern workflows integrate both.
Ecosystem and Open Science Impact
AlphaFold Ecosystem
Large public database
Integration with UniProt
Widely adopted in academia
ESMFold Ecosystem
Open protein language models
Strong integration with large-scale sequence analysis
Growing developer community
The competition between DeepMind and Meta AI has accelerated innovation in computational biology.

Future Outlook
The next wave of protein modeling will likely include:
Better protein complex prediction
Ligand and drug-binding modeling
Multi-state structural predictions
Integration with generative protein design
Multimodal biological AI systems
Both AlphaFold and ESMFold are stepping stones toward fully generative biological design platforms.
AlphaFold and ESMFold represent two complementary approaches to protein structure prediction. AlphaFold emphasizes evolutionary depth and structural refinement. It is the gold standard for high-accuracy research. ESMFold emphasizes scale, speed, and transformer-based intelligence. It excels in rapid, large-scale prediction tasks.
The best choice depends on your priorities. In many advanced workflows, the smartest strategy is not choosing one over the other but combining both. Protein folding is no longer just a biological question. It is now an AI-driven frontier shaping the future of medicine, biotechnology, and scientific discovery.
FAQ's
Q: What is the main difference between AlphaFold and ESMFold?
The core difference lies in how they predict protein structures.
AlphaFold relies heavily on multiple sequence alignments and evolutionary information to achieve very high structural accuracy.
ESMFold uses large protein language models trained on massive sequence datasets and can predict structures directly from a single sequence without requiring MSAs.
In simple terms, AlphaFold focuses on evolutionary depth. ESMFold focuses on language-model intelligence and speed.
Q: Which model is more accurate for protein structure prediction?
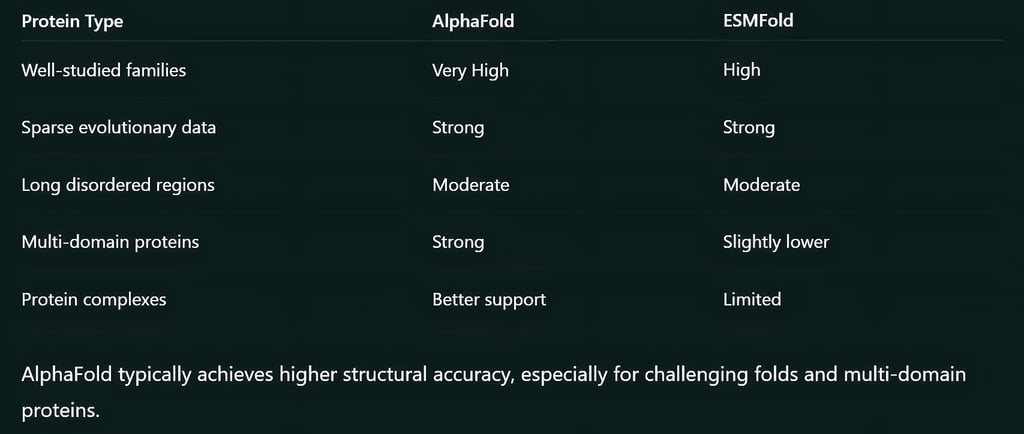
In most benchmarks, AlphaFold achieves slightly higher accuracy, especially for complex or multi-domain proteins.
However, ESMFold performs surprisingly well given its lighter computational pipeline. For many standard proteins, the difference in accuracy is small.
If precision is critical, AlphaFold is usually preferred. If speed and scalability matter more, ESMFold is often the better option.
Q: Is ESMFold faster than AlphaFold?
Yes. ESMFold is significantly faster because it skips the computationally heavy MSA generation step required by AlphaFold.
This makes ESMFold ideal for:
Large-scale protein screening
Metagenomics research
High-throughput pipelines
Q: Do AlphaFold and ESMFold replace laboratory experiments?
No. While both models are highly advanced, they do not fully replace experimental techniques such as X-ray crystallography, cryo-EM, or NMR spectroscopy.
They are powerful prediction tools that:
Guide experiments
Reduce trial and error.
Accelerate hypothesis testing
Experimental validation is still essential for critical applications like drug development.
Q: Can AlphaFold and ESMFold predict protein complexes?
AlphaFold has extended versions that support protein complex prediction with improved performance.
ESMFold primarily focuses on single-chain structure prediction and has more limited support for complex modeling.
For multi-protein interaction studies, AlphaFold-based workflows are generally more mature.
Q: Which model is better for drug discovery?
For structure-based drug design, AlphaFold is often favored because of its slightly higher structural precision and detailed confidence metrics like pLDDT scores.
ESMFold is valuable during early-stage screening when researchers need rapid structural predictions across thousands or millions of sequences.
In practice, many research teams use both tools in complementary workflows.
Q: Are AlphaFold and ESMFold open source?
AlphaFold’s research code and predicted structures are publicly available through its database, though full production pipelines can be complex to run.
ESMFold and its underlying protein language models were released with open research access, making them easier to integrate into scalable AI workflows.
Q: What is the future of AI-based protein folding?
The next stage will likely include:
Improved protein complex prediction
Ligand and small-molecule binding modeling
Dynamic structure prediction
Generative protein design
AlphaFold and ESMFold are foundational systems. Future models will build on their ideas to create more complete biological AI platforms.
Subscribe To Our Newsletter
All © Copyright reserved by Accessible-Learning Hub
| Terms & Conditions
Knowledge is power. Learn with Us. 📚


